┌─ projects ──────────────────────────────────────┐
🦞 Labster Claw
Multi-agent lab management over encrypted group chat
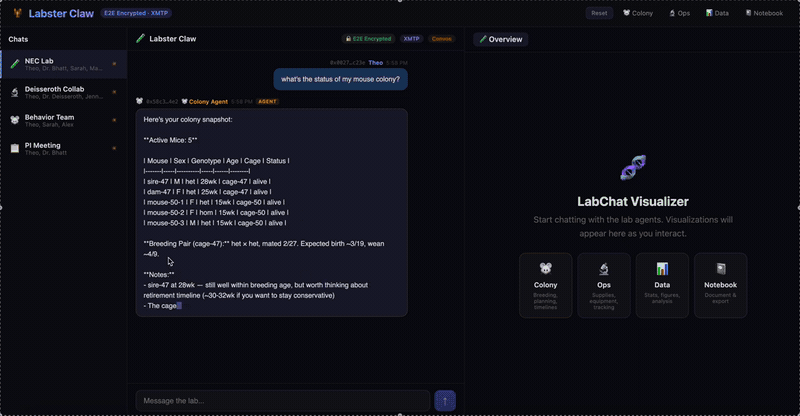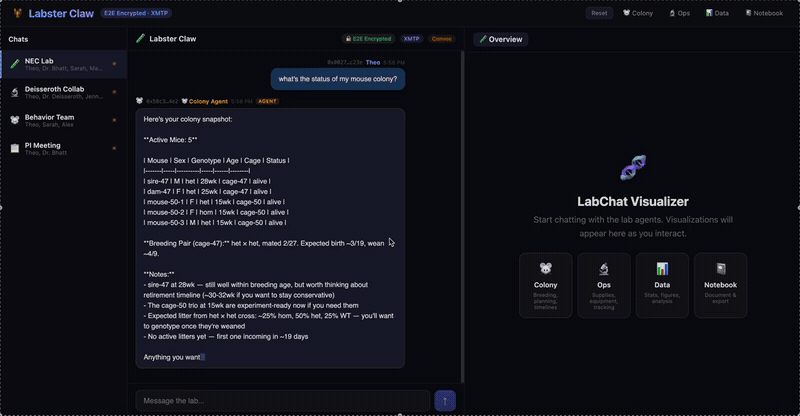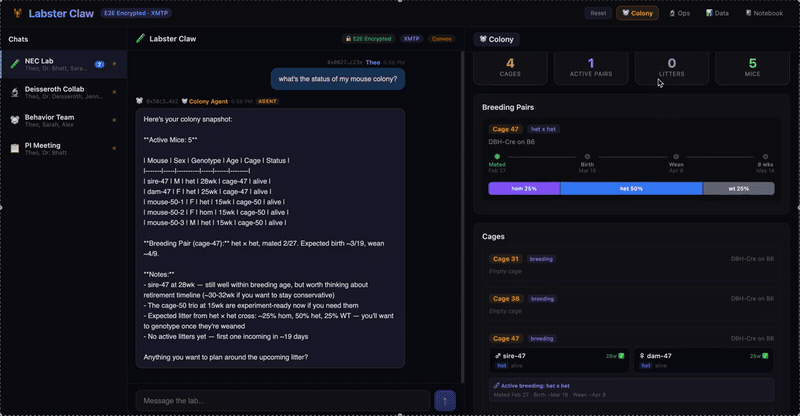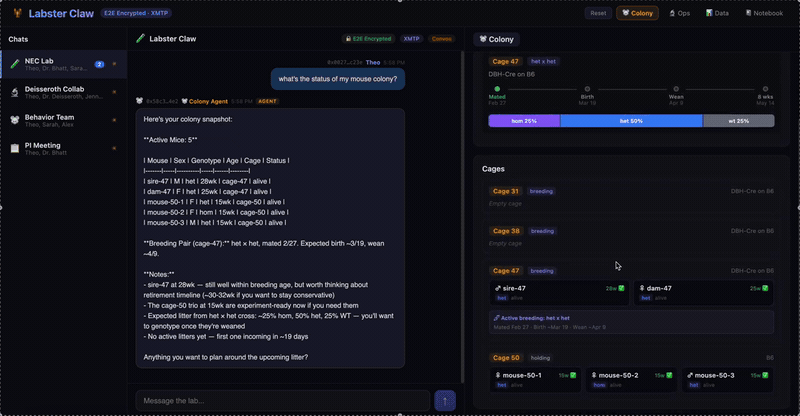
A multi-agent system for neuroscience lab operations built on XMTP encrypted group chat. Three specialized agents — Colony (mouse breeding, JAX schemes, Mendelian genetics), Ops (equipment scheduling, conflict detection), and Data (CSV ingestion, statistical analysis, figure generation) — coordinate through a natural language message router. Lab data is pre-publication and IACUC-sensitive; E2E encryption isn't a feature, it's the architecture.
🧬 Neurochemical Transcriptomics
Brain-wide transcriptomic profiling across neurochemical perturbations
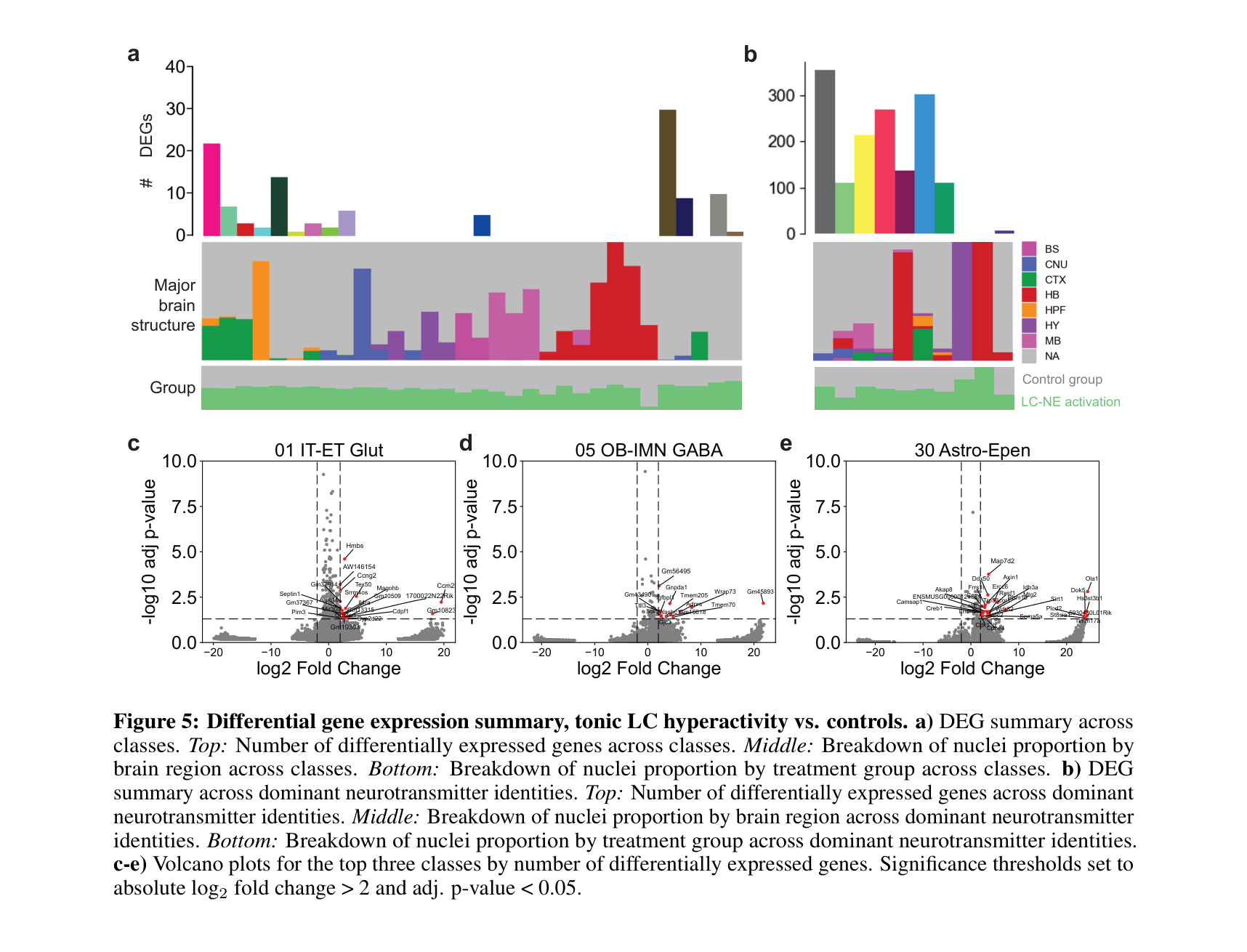
Analysis pipeline for brain-wide single-nucleus RNA-seq across neurochemical perturbations. Includes data preprocessing, dataset integration, cell type composition analysis, differential expression, gene set enrichment (GSEA), and receptor profiling. Built in Python (scanpy/AnnData) and R (Seurat), with reproducible figure generation for the accompanying manuscript.
🧠 Neural Decoders on Area 2 Spiking
Linear, state-space, and RNN decoders on primate somatosensory cortex
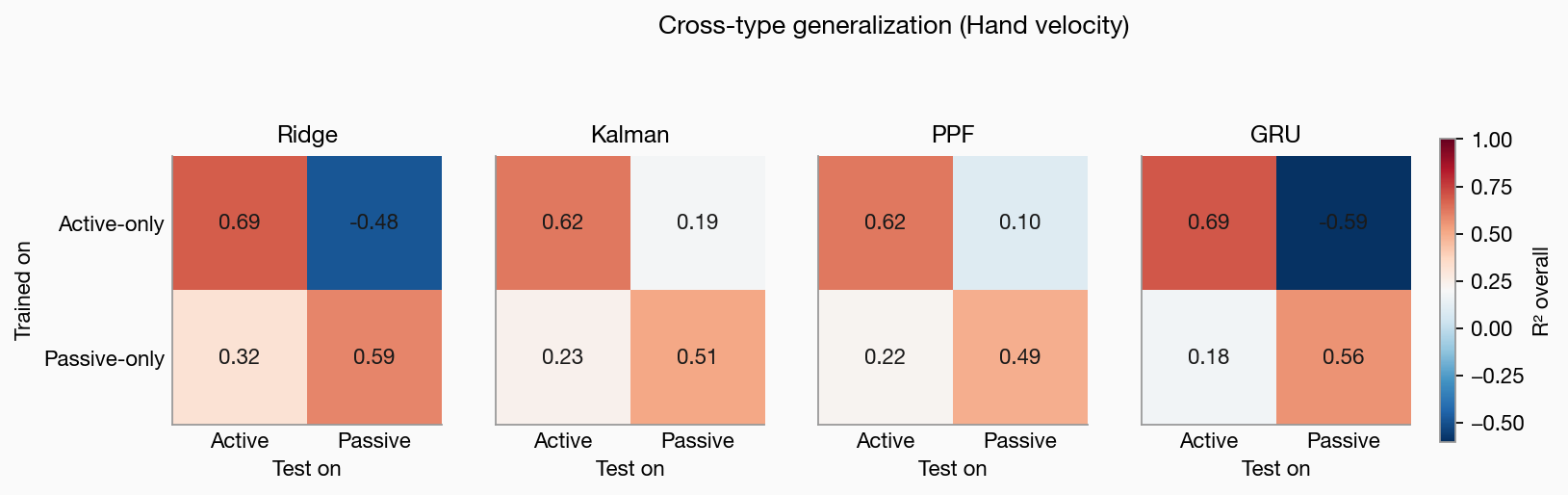
Four decoders — Ridge, Gaussian Kalman (Wu 2006), Point Process Filter (Eden 2004), and a 3-seed GRU ensemble — compared head-to-head on the Neural Latents Benchmark Area2_Bump dataset (primate area-2 spiking during reaching with mechanical perturbations). The interesting finding: when trained only on active reaches and tested on passive bump trials, state-space decoders stay positive (R² ≈ +0.2) while Ridge and GRU collapse to R² ≈ −0.5 — a concrete case study for why an explicit dynamics prior matters under condition shift. Also includes 39-dim muscle-space decoding with a principled state-parameterization choice (pos+vel for hand-space, vel-only for OpenSim-muscle-space) and an honest null result on whether the Poisson likelihood buys you anything over a Gaussian approximation at typical BCI count rates.
🔬 Histology ROI Quantification & Colocalization
Fluorescence colocalization analysis with semi-automated LC region detection
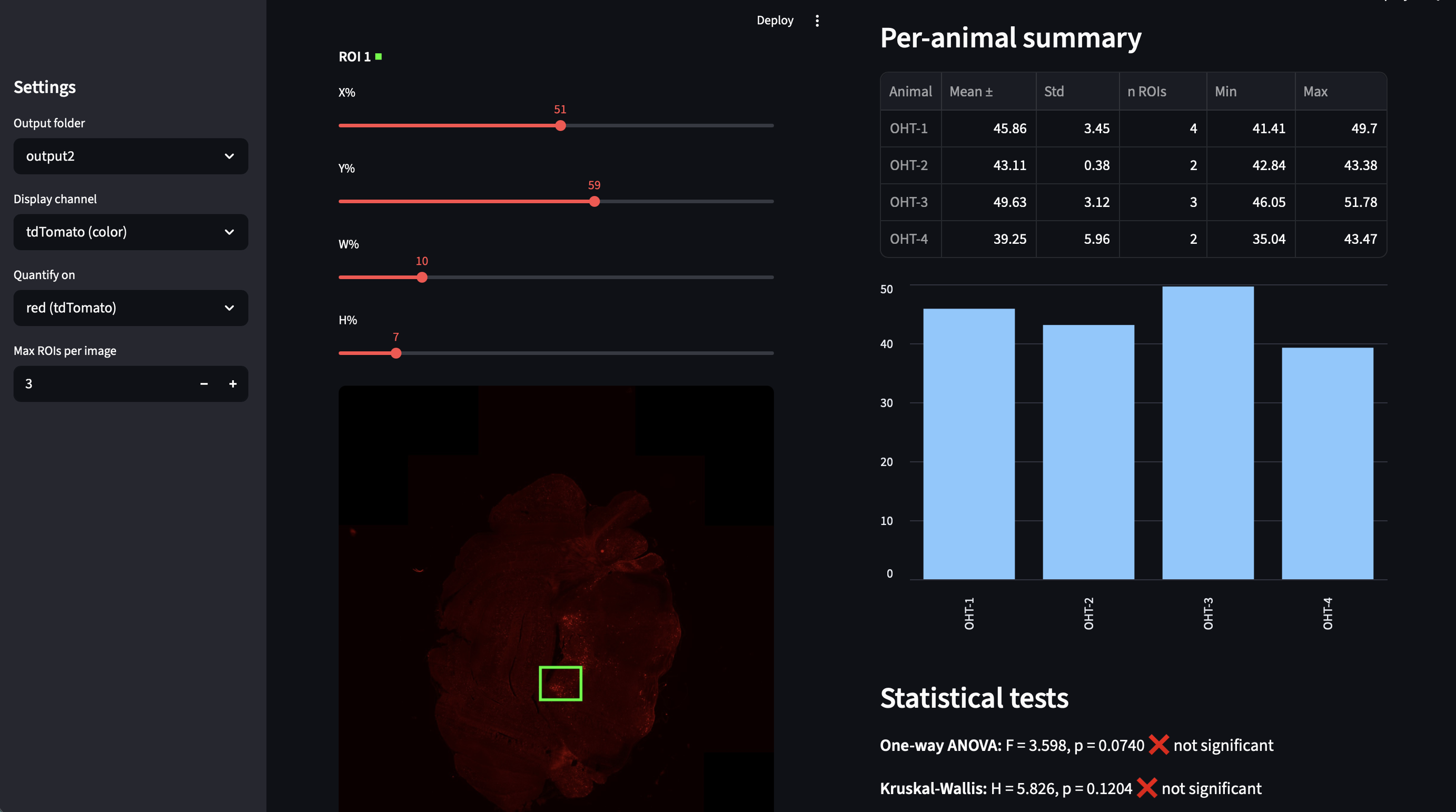
A pipeline for processing 3-channel .nd2 fluorescence microscopy images (TH/tdTomato/DAPI) with globally consistent contrast normalization, region-level DAPI normalization, and TH-tdTomato colocalization analysis in the locus coeruleus. Includes semi-automated LC detection that ranks TH+ clusters by intensity for manual ROI selection, restricting analysis to verified LC regions. Generates DAPI-normalized contact sheets with green/yellow TH+ signal highlighted against grayscale backgrounds for visual QC. Supports whole-section and LC-restricted colocalization, TH+ expression sex-difference analysis, interactive Streamlit ROI quantification, and publication-ready plots with significance bars. Built for TRAP2-Ai14 histology.
📊 GLM-HMM Comparison Tool
Cross-validated behavioral state model fitting and comparison
A toolkit for fitting and comparing Generalized Linear Model Hidden Markov Models (GLM-HMMs) in behavioral neuroscience. Implements cross-validated model selection across state counts (K=1–4) for both standard and restructured architectures, addressing circularity concerns when modulatory variables (e.g., arousal) are used to both define states and analyze within-state dynamics. Includes a CLI for global fitting with per-trial state assignment extraction, posterior probabilities, and parameter export. Based on the Ashwood et al. (2022) framework.
└─────────────────────────────────────────────────┘