┌─ work ──────────────────────────────────────────┐
├─ writing ───────────────────────────────────────┤
The Institutional Lag
Post-labor economics, AI displacement, and the design of antifragile institutions.
[ writing... ]Our Genetic Constitution
On the biological foundations of political order and why genetic engineering will force us to rethink the social contract.
The Case for Human Hibernation
Deep space travel, longevity, and the engineering challenge of putting humans into suspended animation.
├─ projects ──────────────────────────────────────┤
🦞 Labster Claw
Multi-agent lab management over encrypted group chat
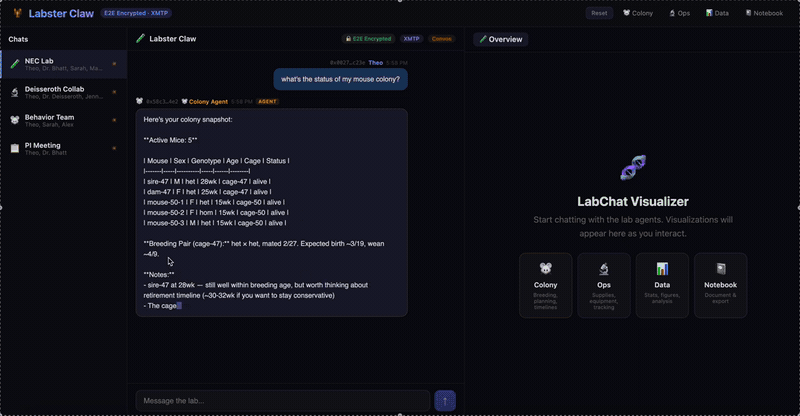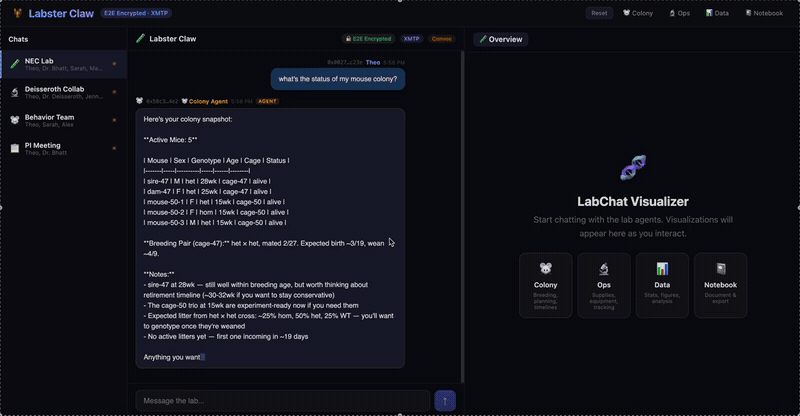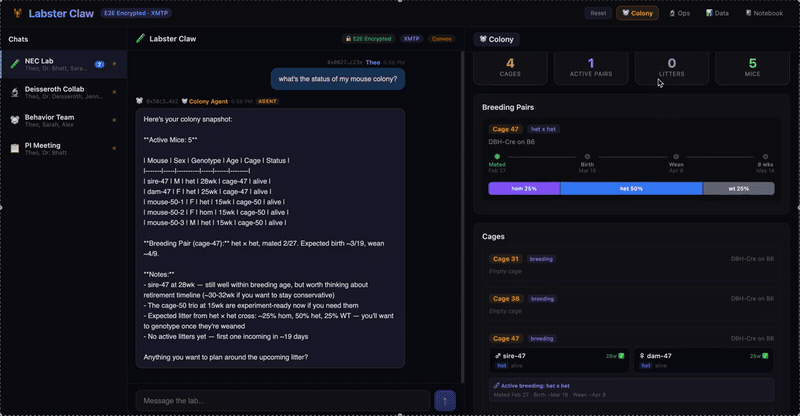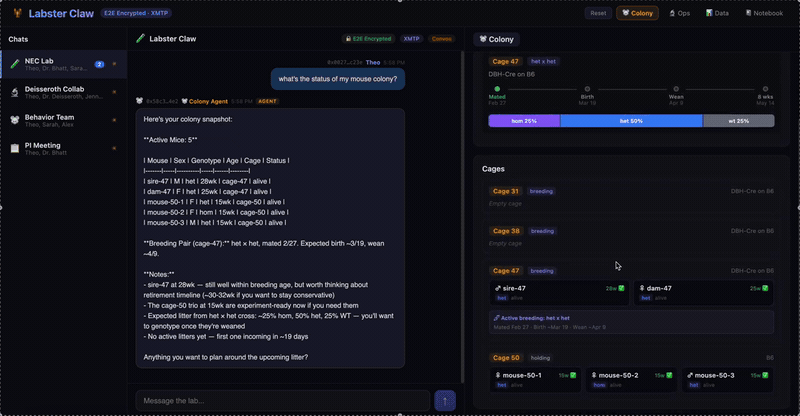
A multi-agent system for neuroscience lab operations built on XMTP encrypted group chat. Three specialized agents — Colony (mouse breeding, JAX schemes, Mendelian genetics), Ops (equipment scheduling, conflict detection), and Data (CSV ingestion, statistical analysis, figure generation) — coordinate through a natural language message router. Lab data is pre-publication and IACUC-sensitive; E2E encryption isn't a feature, it's the architecture.
🧬 Neurochemical Transcriptomics
Brain-wide transcriptomic profiling across neurochemical perturbations
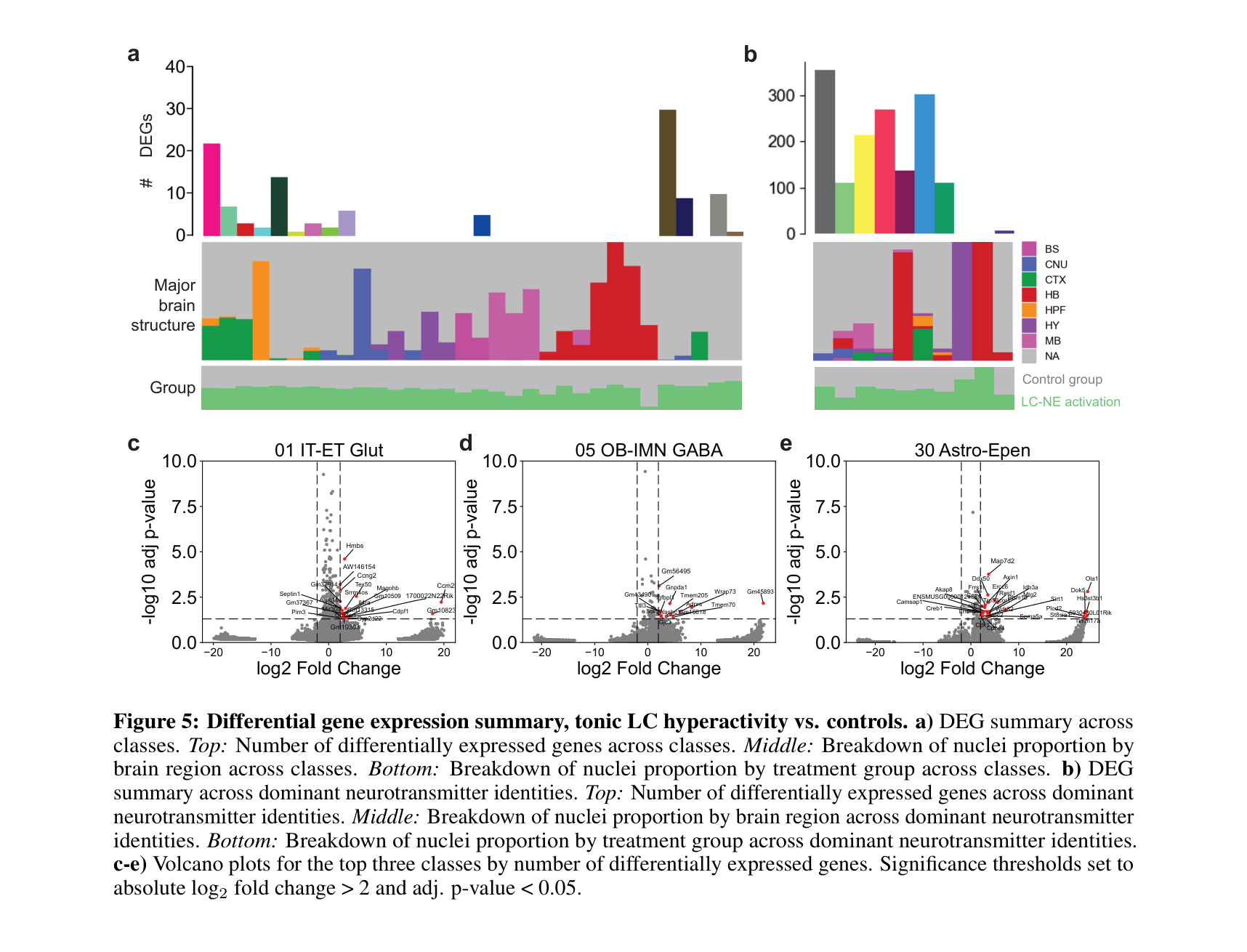
Analysis pipeline for brain-wide single-nucleus RNA-seq across neurochemical perturbations. Includes data preprocessing, dataset integration, cell type composition analysis, differential expression, gene set enrichment (GSEA), and receptor profiling. Built in Python (scanpy/AnnData) and R (Seurat), with reproducible figure generation for the accompanying manuscript.
🔬 Histology ROI Quantification
Standardized contrast normalization and interactive fluorescence quantification
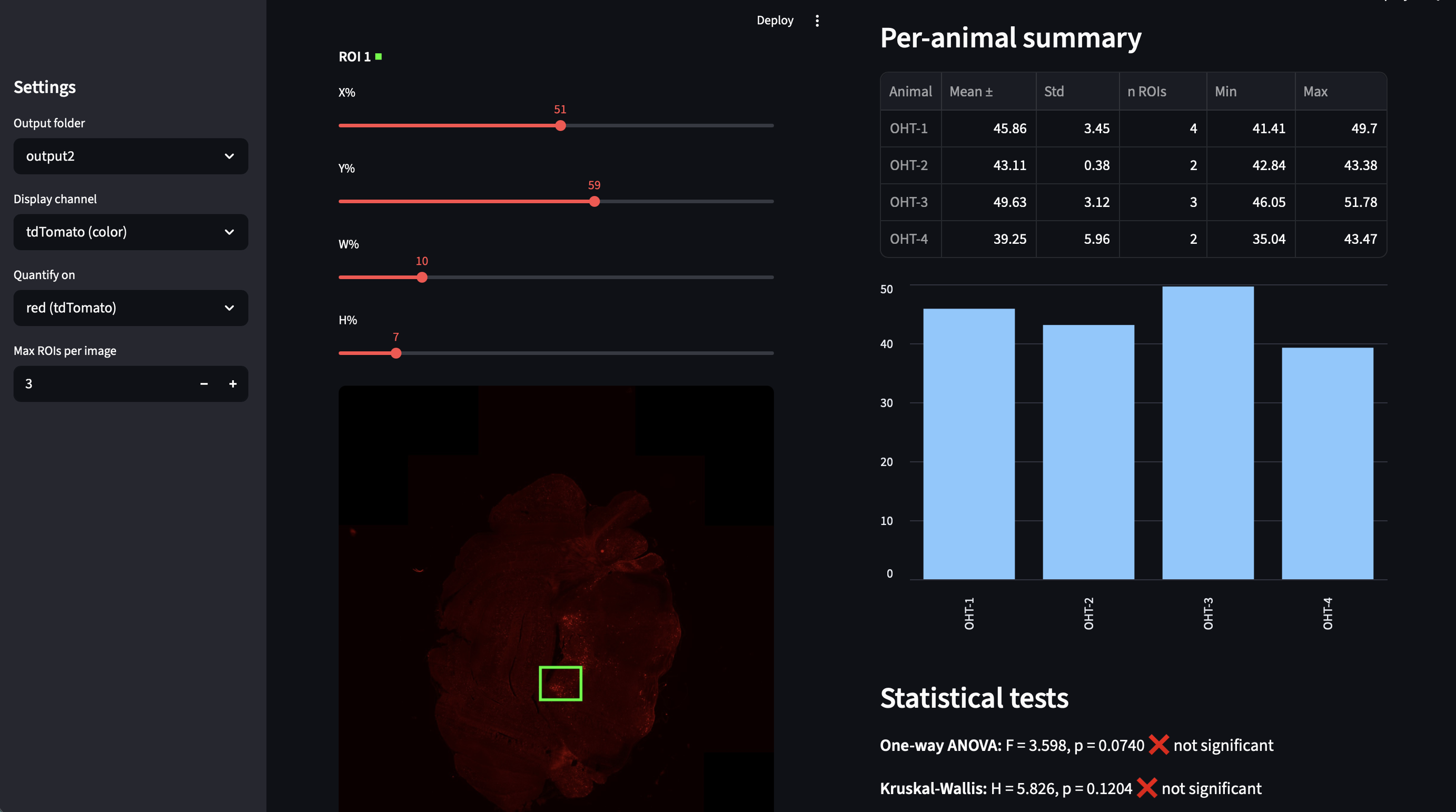
A pipeline for processing .nd2 fluorescence microscopy images with globally consistent contrast normalization, plus an interactive Streamlit app for ROI-based expression quantification across animals. Two-pass processing computes global percentile-based intensity limits across all images, then applies uniform scaling. The web app supports multiple ROIs per image, multi-image selection per animal, automatic ANOVA and pairwise statistics, and CSV export. Built for TRAP2-Ai14 (tdTomato/DAPI) histology but generalizable to any multi-channel fluorescence data.
📊 GLM-HMM Comparison Tool
Cross-validated behavioral state model fitting and comparison
A toolkit for fitting and comparing Generalized Linear Model Hidden Markov Models (GLM-HMMs) in behavioral neuroscience. Implements cross-validated model selection across state counts (K=1–4) for both standard and restructured architectures, addressing circularity concerns when modulatory variables (e.g., arousal) are used to both define states and analyze within-state dynamics. Includes a CLI for global fitting with per-trial state assignment extraction, posterior probabilities, and parameter export. Based on the Ashwood et al. (2022) framework.
└─────────────────────────────────────────────────┘